The alphabet of life
The grand tale of life is long and complicated. Storylines intertwine and many subplots twist and turn unexpectedly. Amazingly, this billion-year-spanning story is written in an alphabet that contains only four letters, the alphabet of DNA. A for adenine, C for cytosine, G for guanine, and T for thymine. That’s it. That’s all that’s needed to compose the paragraphs and chapters in the book of life.
To understand how this works we need to travel to the inside of the building blocks that make living organisms, their cells. From the tiniest bacterium to the mightiest whale, every living thing carries within itself an immense library that contains all the information necessary to build a fully functional organism. This library is shaped like a ladder. Two rails of alternating sugars and phosphates are connected by rungs. Each half of such a rung is constructed out of a base that functions as a letter in life’s alphabet. And these letters are quite picky. A only connects to T, while C’s preferred partner is G. More flexible than any real ladder, the rails coil around each other and form the famous double helix of DNA.

This double helix, in turn, twists and curls around itself, compressing the entire library until it is small enough to be stuffed into a single cell. In so-called prokaryotes, a group that includes our bacterium, this collection of DNA floats around freely in the cell. In eukaryotes, including the whale and ourselves, most of the DNA is stuffed into the nucleus, a compartment that can be found in most cells.
The entire library is so expertly packed that it is stored in a space too tiny to observe without powerful microscopes. It gets even more remarkable when you add the fact that, depending on the organism, the collection of DNA contains several thousands to tens of billions of A’s, C’s, G’s, and T’s.
Adding letters
When you look at the marvels of the living world all around you, it seems incredible that all those stories of birth and death, of life and love, are written with an alphabet just four letters long. How is this even possible? In the years following the discovery of DNA’s double helix structure in 1953, some of the most impressive minds in biology devoted themselves to exactly this question. Eventually, they uncovered a deceptively simple system that allows this short alphabet to give rise to diverse and complicated sentences.
In short, every combination of three DNA letters 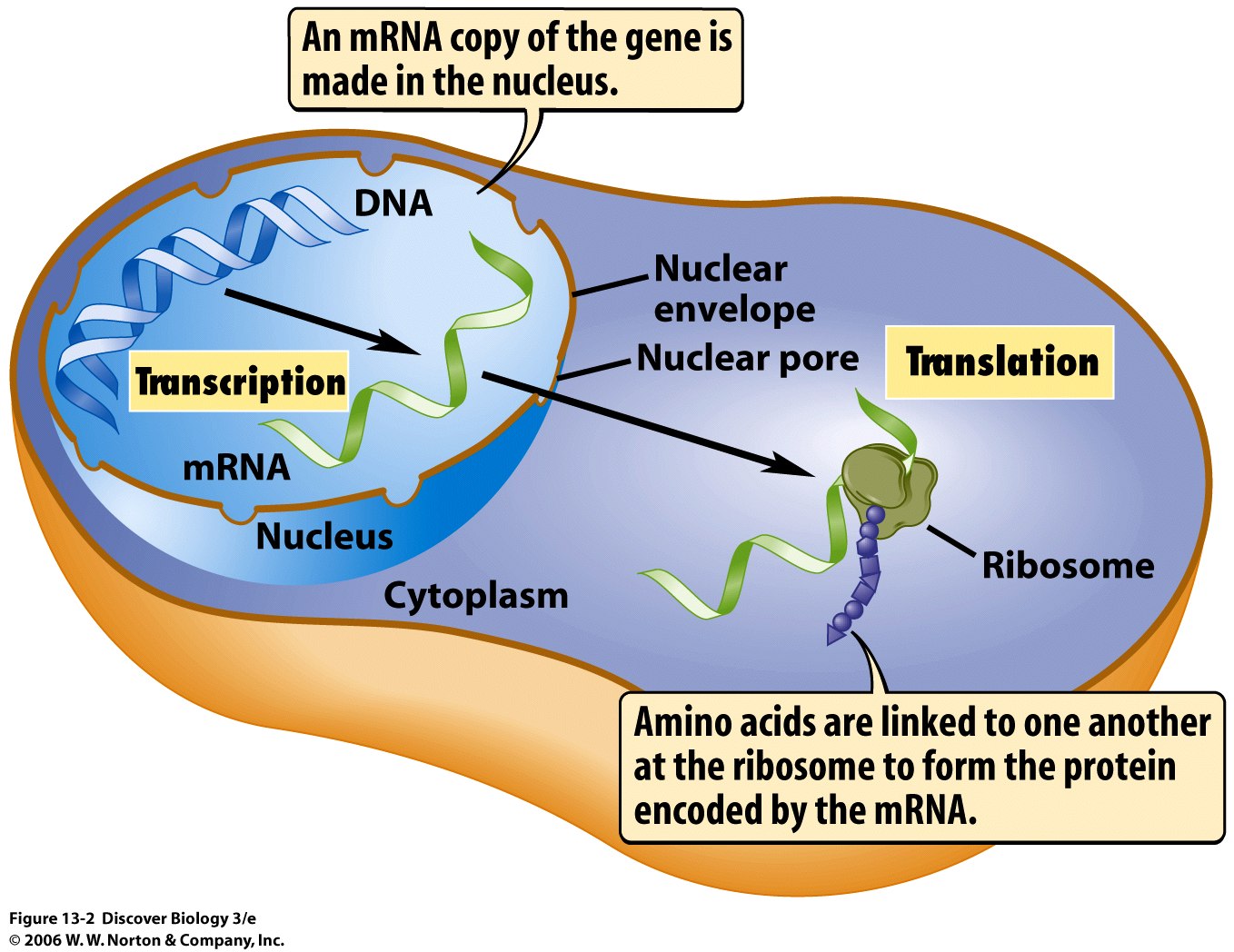 (known as a ‘codon’) codes for a compound known as an amino acid. These amino acids are strung together into complex 3D shapes to form proteins, which are molecules that perform a lot of important tasks in an organism’s body. This decoding from DNA to protein requires transcription into an intermediate template, known as RNA, rendering the DNA code into a quite similar four letter language (only T is replaced by U for uracil). The RNA can then be translated by the protein-making machinery in the cell and this completes the decoding process.
(known as a ‘codon’) codes for a compound known as an amino acid. These amino acids are strung together into complex 3D shapes to form proteins, which are molecules that perform a lot of important tasks in an organism’s body. This decoding from DNA to protein requires transcription into an intermediate template, known as RNA, rendering the DNA code into a quite similar four letter language (only T is replaced by U for uracil). The RNA can then be translated by the protein-making machinery in the cell and this completes the decoding process.
Each letter in a codon can be A, C, G or T, so there are (4 x 4 x 4 =) 64 possible three-letter combinations of DNA bases that code for 20 amino acids (many amino acids have more than one corresponding DNA codon and there are a few ‘stop’ signals as well). Since most proteins are made out of dozens to hundreds, and sometimes even tens of thousands, amino acids, the number of combinations is quite large. All this based on a four-letter alphabet.
But what if we could add letters? What if we could extend the alphabet of life?
Spurred on by these questions, several groups of biochemists and molecular biologists set themselves to the task of designing new letters. Time to make our own additions to the story of life.
Writing new paragraphs
Of course, concocting new DNA letters is only the first step. If the additions are meaningless, or worse, distort the meaning of the rest of the alphabet, no novel paragraphs will be written. The new letters have to integrate into the already existing DNA. In other words, true success lies in the capacity of the man-made letters to be copied and translated alongside their older fellows.
This has proven to be a challenge. But, in the past few years, several teams have managed to overcome it. In fact, some expanded and functional six-letter alphabets have already been put together. Whether the new letters are called P and Z, or carry more complex names such as 5SICS and NaM, these extended alphabets have been shown to be copied and translated just as good as the basic one. And what’s more, they do not undergo these processes flawlessly. They can actually mutate just like regular DNA, allowing evolution to take its course using six letters instead of the usual four.
So, we have expanded alphabets, and they work. But can we use them to contribute to life’s grand tale? It turns out we can. The team behind 5SICS and NaM has succeeded in integrating their new letters into the DNA of a bacterium known as Escherichia coli, a popular organism in a lot of biological research. Not only did this little critter accept the new letters, but it replicated, transcribed, and translated them just like the A’s, C’s, G’s and T’s. It effectively became the first known living organism with a six-letter alphabet making up its vital biological library.
These initial story additions that use newly designed letters are still short and limited, but they open up a vast space of possible plot twists to be written as we get more proficient in using extended DNA alphabets. The content of these storylines could be as multifarious as life itself. Initial ideas involve thrillers about applied biosafety, adventures into realms of unknown or improved proteins, and chronicles from the clinic about detecting and monitoring diseases.
As progress is made on the implementation of new letters in life’s alphabet, the only limit seems to be our own imagination. Dystopian dread, utopian euphemisms or something more realistic altogether, what will it be? To successfully add paragraphs to a great story, careful copy-editing, reader feedback, collaborative writing, and the recognition of multiple interpretations can all help. Especially when using the language of life.
Whatever new plots that will emerge, they’ll surely be exciting.
References:
Hirao I, & Kimoto M (2012). Unnatural base pair systems toward the expansion of the genetic alphabet in the central dogma. Proceedings of the Japan Academy. Series B, Physical and biological sciences, 88 (7), 345-67 PMID: 22850726
Yang, Z., Chen, F., Alvarado, J., & Benner, S. (2011). Amplification, Mutation, and Sequencing of a Six-Letter Synthetic Genetic System Journal of the American Chemical Society, 133 (38), 15105-15112 DOI: 10.1021/ja204910n~
Malyshev, D., Dhami, K., Quach, H., Lavergne, T., Ordoukhanian, P., Torkamani, A., & Romesberg, F. (2012). Efficient and sequence-independent replication of DNA containing a third base pair establishes a functional six-letter genetic alphabet Proceedings of the National Academy of Sciences, 109 (30), 12005-12010 DOI: 10.1073/pnas.1205176109
Malyshev, D., Dhami, K., Lavergne, T., Chen, T., Dai, N., Foster, J., Corrêa, I., & Romesberg, F. (2014). A semi-synthetic organism with an expanded genetic alphabet Nature, 509 (7500), 385-388 DOI: 10.1038/nature13314
Image credits 1: Magnus Manske: Location of eukaryote nuclear DNA within the chromosomes.
Image credits 2: csus.edu





