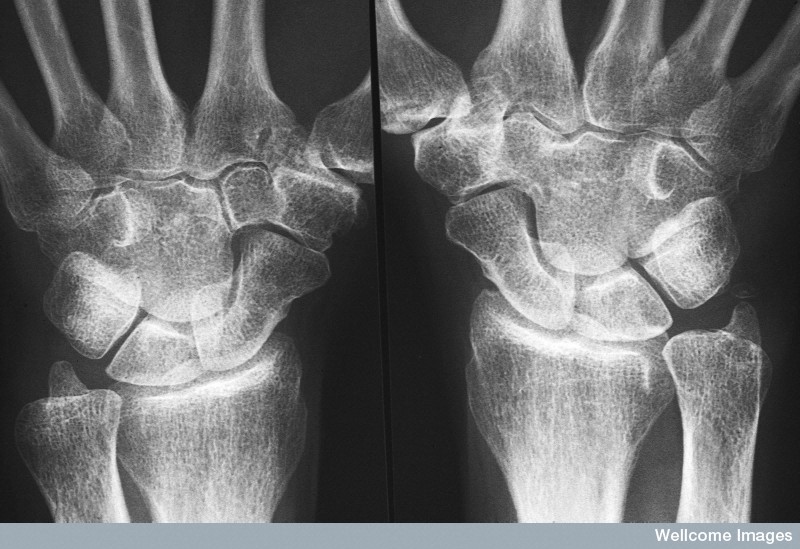
A disease of unknown causes
Rheumatoid arthritis (RA) is an auto-immune disease which primarily leads to inflammation of the joints. The exact cause of this chronic illness remains unknown, yet genetic factors and environmental exposes, such as smoking, are known to play a role in the development of the disease.
RA-patients experience severe pain and their affected joints are abnormally swollen. Eventually, the prolonged inflammation leads to irreversible joint damage and invalidity.

management. Source: Smolen, 2017 [1]
The primary goal of the anti-inflammatory treatment is, therefore, to rapidly reduce disease activity, which will improve both short term and long term patients’ outcomes. With a vast arsenal of treatments, among which [su_tooltip style=”cluetip” position=”north” rounded=”yes” size=”2″ content=”TNFs’s reduce inflammation by targeting the inflammation-causing substance called Tumor Necrosis Factor (TNF)” close=”yes”]tumor necrosis factor-alpha inhibitors (TNFi’s)[/su_tooltip], RA disease activity decreases sufficiently in most but not all cases.
As non-responders to treatment are at risk of worse outcomes, the identification of non-responders before initiation of therapy would aid in making strategic treatment decisions, for instance starting different treatment with better chances of a response. Biomarker research in this field has aimed to find such predictive factors or ‘predictors’ of response. In this research, new potential targets that predict response have frequently been identified, yet validation across multiple cohorts has failed so far.
Predicting response to treatment
In this research, data from more than 400 RA patients, from the “Biological and Outcome Compared and predicted Utrecht region in Rheumatoid Arthritis” (BiOCURA) study, were used. Within serum and isolated blood cells, we searched for potential biomarkers to predict the response to biological treatment in RA, by screening DNA, mRNA, microRNA, proteins and metabolites.

Derivation of biomarkers. DNA is copied into RNA, which is consecutively translated to proteins. This process is influenced by epigenetic mechanisms. Proteins are responsible for biochemical processes, leading to metabolite formation and conversion. Images of biomarker structures were derived from Wikipedia.
The lack of biological predictors
Still, after an extensive search, no predictors of response to RA treatment are available ready to use in clinical practice. The lack of identified predictors does not exclude the possibility of prediction of response, as many reasons for the inability to find biologically relevant predictors can co-exist. It is, therefore, conceivable that an adoption of the strategy enables the field to find reproducible and relevant predictors.
However, the possibility exists that prediction of response is too difficult in practice. Either the biology of RA might be too complex and variable to get hold of or the patients’ response to unknown parameters, such as the exposome, may be very unpredictable, or both. Still, preeminent scientists in rheumatology firmly believe that predictors for treatment response in RA will be available someday to use.
In a nutshell
To get there, we need a better understanding of the pathogenesis and pharmacological effects of therapy. Instead of the current ‘shotgun approach’ in which many biomarkers at once are measured, and some might hit the target, energy and funding might better be spent in understanding the pathophysiology and pharmacological response mechanisms. Using this knowledge, hypotheses can be made that can be individually tested, while also taking into account all (biologically possible) influencing factors. Although this might seem a step backward, it can actually help us forward.
An adjustment in the study design is necessary to pick up robust predictors of response to therapy in RA. Efforts should be made to replicate findings after biomarker discovery and publish all validated and unvalidated results.
The measurement of response in RA is a major challenge and should be further improved to prevent misclassification, for instance by implementing pure biochemical response measurement. The high degree of heterogeneity in clinical characteristics among RA patients should be used to correct for the substantial effect on biomarkers levels, which will increase the generalizability of results. New data analyses techniques, such as systems biology, should be further developed. However, a better understanding of the pathogenesis and pharmacological effects of therapy is needed most.
References:
[1] Smolen, J., & et al. (2017). EULAR recommendations for the management of rheumatoid arthritis with synthetic and biological disease-modifying antirheumatic drugs Annals of the Rheumatic Diseases, 69 (6), 964-975 DOI: 10.1136/ard.2009.126532
Read the full doctoral research here. [Open Access]
[su_box title=”Share your Science” style=”glass” title_color=”#ffffff”] The Share your Science section aims at giving students and researchers a space to share their thesis to a broader audience. UA Magazine wants to increase the visibility of recent academic work and serve as a bridge between universities and societies. [/su_box]